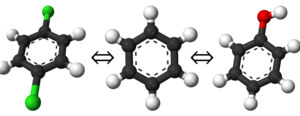
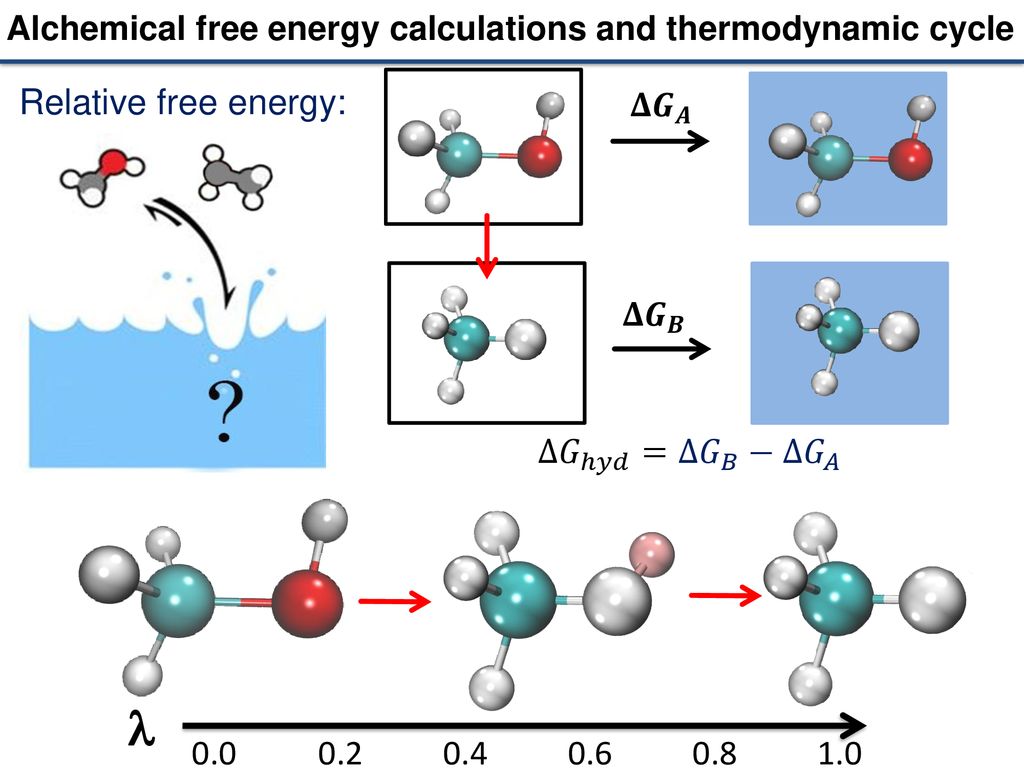
Alchemical Free Energy Calculations in Biomolecules – BioExcel – Centre of Excellence for Computation Biomolecular Research

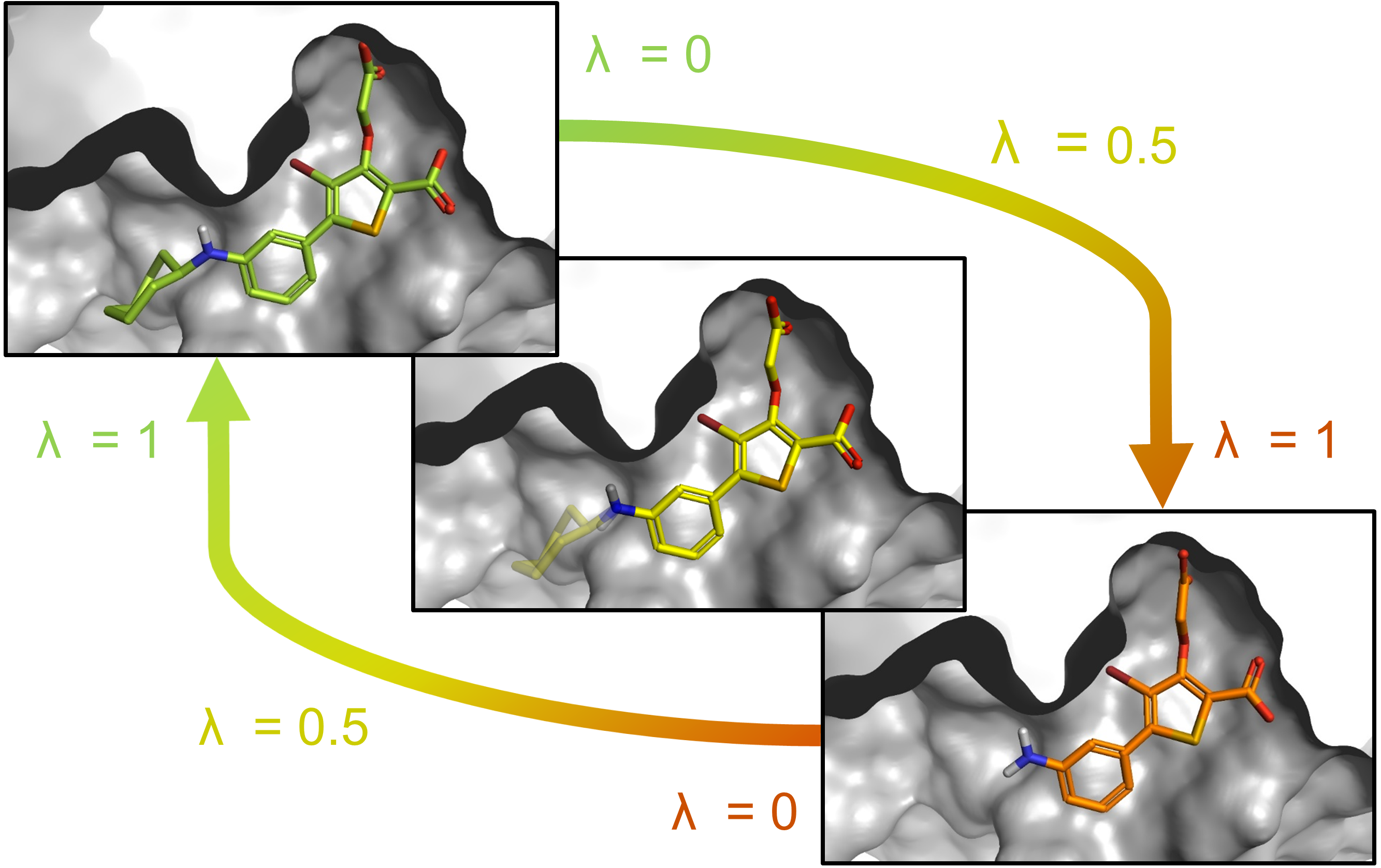
Figure 1 | Absolute Alchemical Free Energy Calculations for Ligand Binding: A Beginner's Guide | SpringerLink

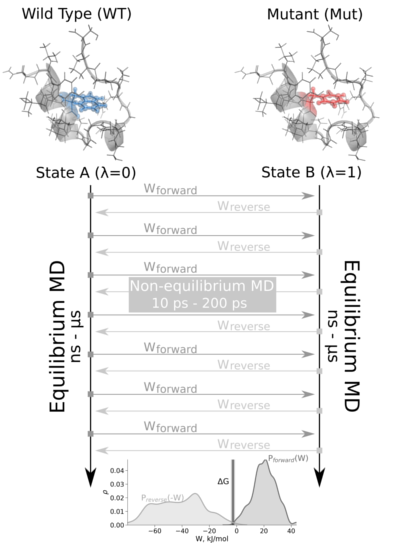
Biomedicines | Free Full-Text | High-Throughput Molecular Dynamics-Based Alchemical Free Energy Calculations for Predicting the Binding Free Energy Change Associated with the Selected Omicron Mutations in the Spike Receptor-Binding Domain of SARS-CoV-2
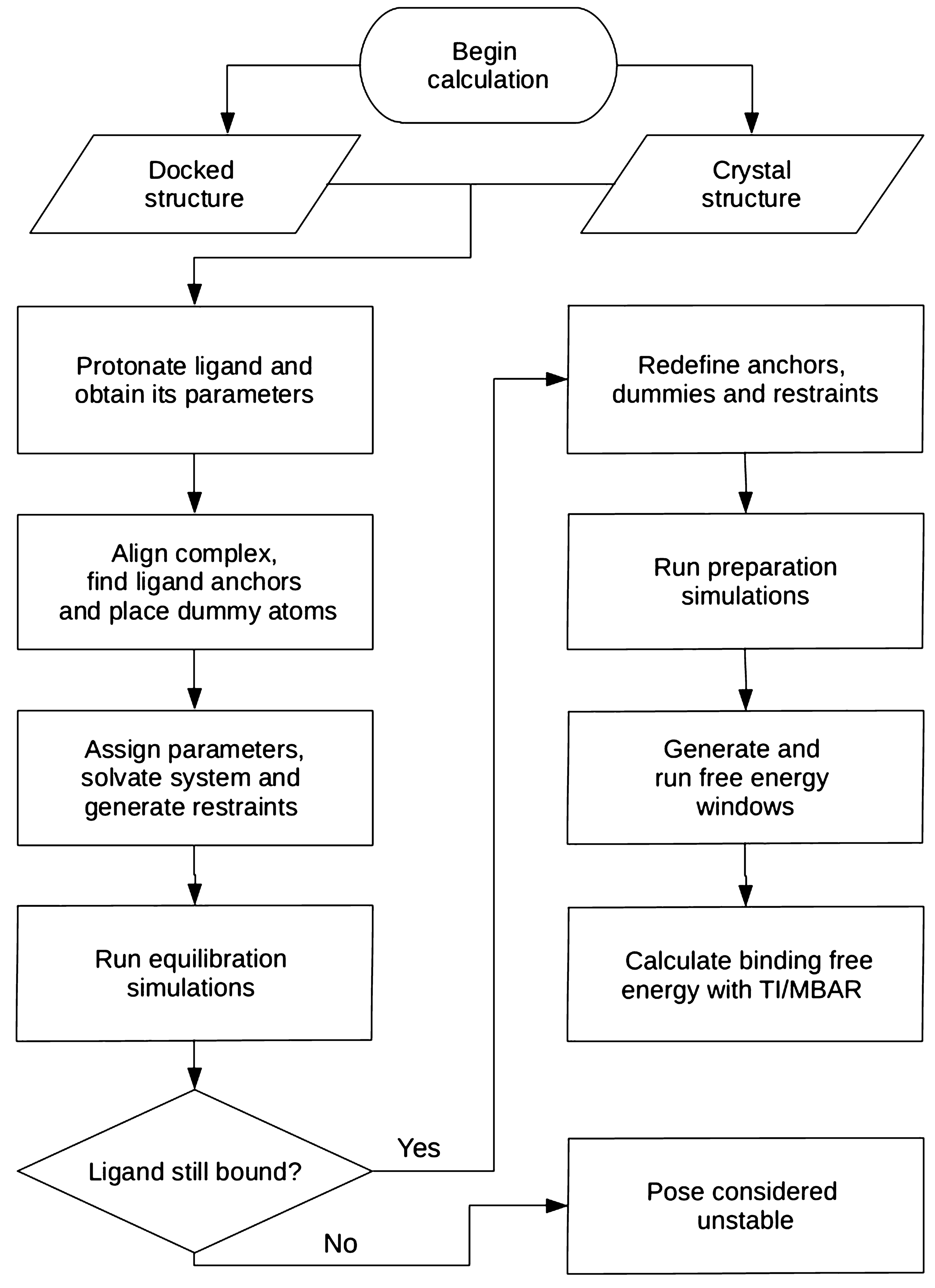
Automation of absolute protein-ligand binding free energy calculations for docking refinement and compound evaluation | Scientific Reports

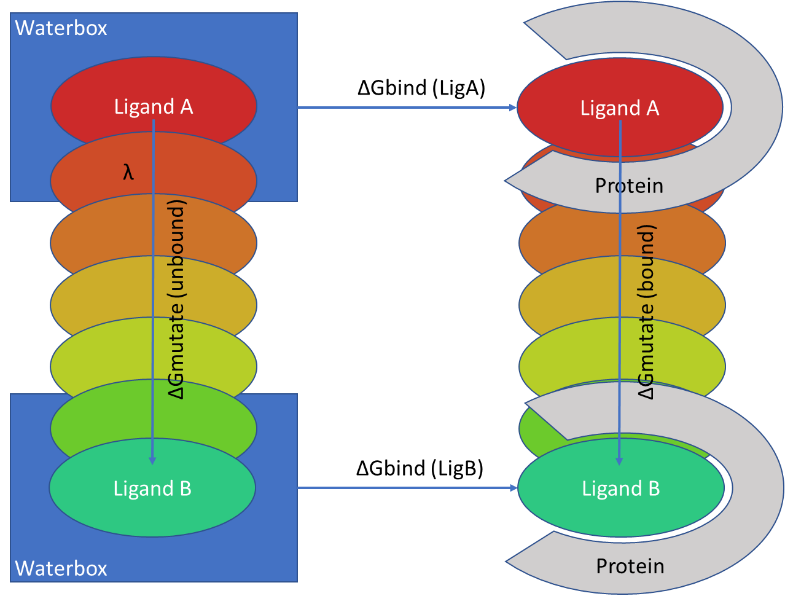
A diagram of how input for relative alchemical free energy calculations... | Download Scientific Diagram
![Best Practices for Alchemical Free Energy Calculations [Article v1.0] | Living Journal of Computational Molecular Science Best Practices for Alchemical Free Energy Calculations [Article v1.0] | Living Journal of Computational Molecular Science](https://livecomsjournal.org/public/journals/15/article_1113_cover_en_US.jpg)
Best Practices for Alchemical Free Energy Calculations [Article v1.0] | Living Journal of Computational Molecular Science

The pmx Webserver: a Powerful Tool for Free Energy Calculations – BioExcel – Centre of Excellence for Computation Biomolecular Research

Alchemical Free Energy Calculations to Investigate Protein–Protein Interactions: the Case of the CDC42/PAK1 Complex | Journal of Chemical Information and Modeling

John Chodera (he/him) on Twitter: "Alchemical free energy calculations can make mechanism-based predictions of many properties relevant to drug discovery, including affinity, selectivity, lipophilicity, resistance, and thermostability, with great ...

Alchemical free energy calculations via metadynamics: Application to the theophylline‐RNA aptamer complex - Tanida - 2020 - Journal of Computational Chemistry - Wiley Online Library

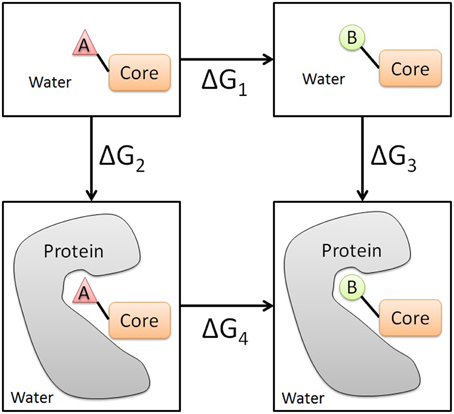
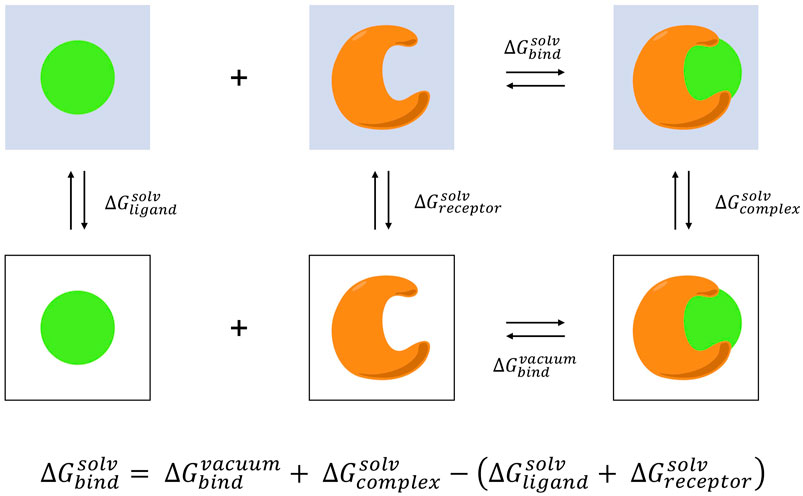
Thermodynamic cycle required for an absolute free energy calculation... | Download Scientific Diagram

Alchemical Hydration Free-Energy Calculations Using Molecular Dynamics with Explicit Polarization and Induced Polarity Decoupling: An On–the–Fly Polarization Approach | Journal of Chemical Theory and Computation
Thermodynamic cycle required for an absolute free energy calculation... | Download Scientific Diagram

Predicting resistance of clinical Abl mutations to targeted kinase inhibitors using alchemical free-energy calculations | Communications Biology
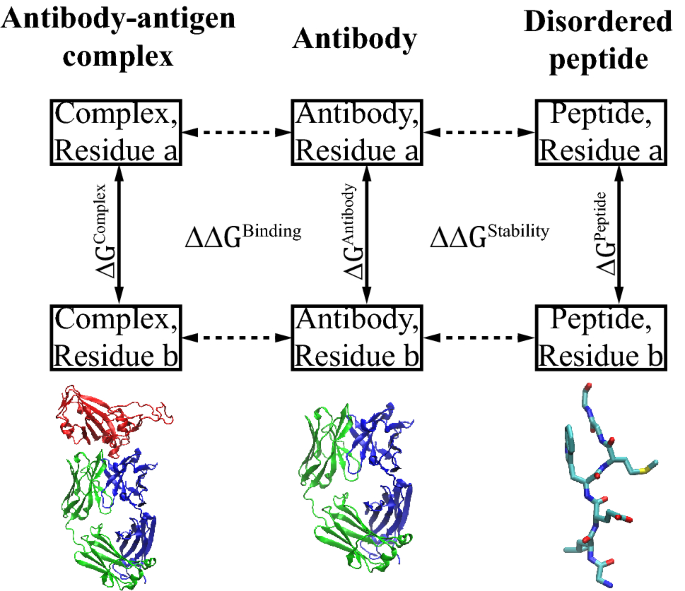
Large-scale application of free energy perturbation calculations for antibody design | Scientific Reports